 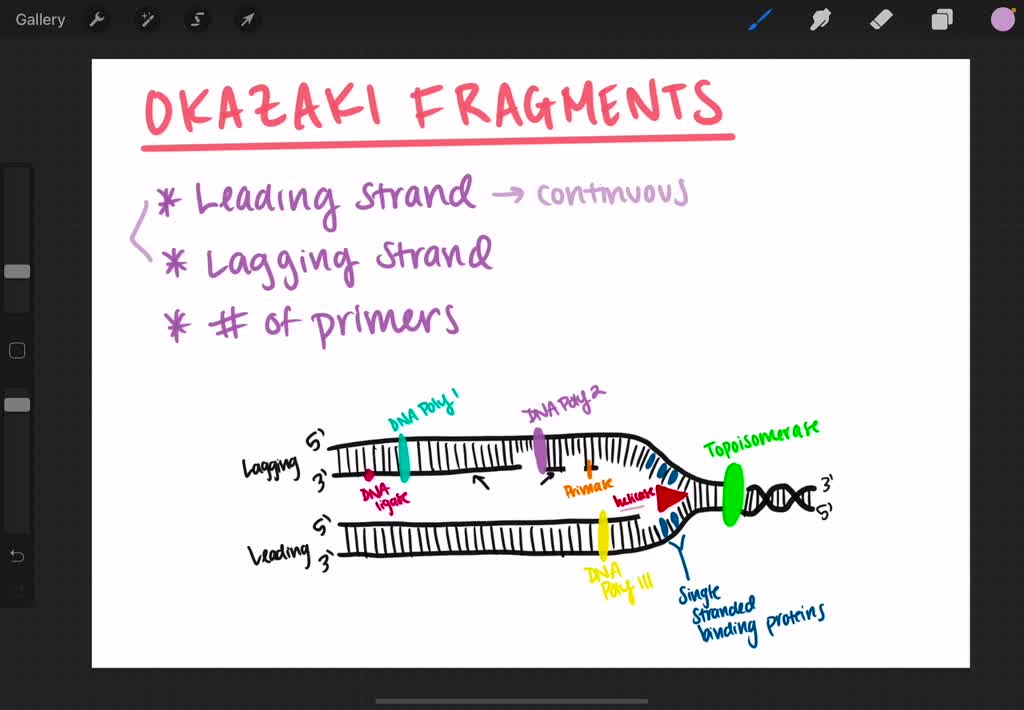
Because completed Okazaki fragments (short nascent chains) wereisolated before the analysis, the labeled nucleotides incorporated at theearliest times had to be added as part of the process of completing themolecule. 11 (b) The nascent DNA chain grows in a 5’ to 3’direction. The 8 to 10S DNA consists of Okazaki fragments, and the fastsedimenting, large DNA contains the growing chains after the Okazaki fragmentshave been joined to them.Īnswer 5. This is consistent with discontinuous DNA synthesis of thelagging strand. As the pulse time increases,more of the labeled thymidine or thymine appears in large DNA, sedimenting atgreated than 60S. Notethat this would generate an equal number of positive superhelical turns, iftopoisomerases were not acting as a swivel during replication.ī)False, they are formed during synthesis of the lagging strand of DNA.Īnswer 5.11 (a) Atshort pulse times (5 sec), the labeled thymidine or thymine appears exlusivelyin small DNA chains sedimenting at about 8 to 10S. Thenumber of helical turns = number of base pairs/number of base pairs per helicalturn. During DNA replication, thecomplementary strands of DNA must unwind completely to allow the synthesis of anew strand on each template. Thenumber of base pairs per helical turn for B-DNA is about 10. The parental DNA remains heavy (HH) but is diluted out by the progenyDNA, which is light (LL). 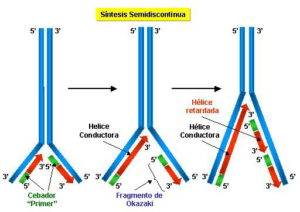
Since no hybrid density DNA is formed, replication is notsemi-conservative or distributive. By moving in this direction after initially binding tothe single-stranded DNA, it will encounter the duplex including molecule B, andthen displace it by its helicase activity. PriAtracks alongthe single-stranded DNA in a 3' to 5' direction (relative to thesingle-stranded DNA). coli chromosome is replicated from one origin, called oriC. For bidirectional replication, thisrequires only one origin, and indeed this is the case. Dividing the size of the chromosome by this amountsynthesized per fork gives 4.64 ´ 10 6bp / 2.4 ´ 10 6 bp, or 1.93.Hence two replication forks are sufficient. Thus in 40 min,one replication fork replicates 60,000 bp per min ´ 40 min = 2.4 ♁0 6 bp. As the replication fork moves 60,000 nucleotidesper min, it produces both daughter strands at the same rate. The reverse of the polymerizationreaction is pyrophosphorolysis (adding a molecule of pyrophosphate across thebond that is broken), resulting in a nucleoside tri phosphate and a DNA chain shorter by one nucleotide.Īnswer 5.6. Removal of a nucleotide from the 3’ end of thegrowing chain by a 3’ to 5’ exonuclease catalyzes the hydrolysis of the 3’nucleotide (adding a molecule of water across the bond that is broken),generating a nucleoside mono phosphateand a DNA chain shorter by one nucleotide. Note thatif an incorrectly incorporated nucleotide were removed by a proofreadingexonuclease (a 5’ to 3’ exonuclease in this hypothetical example), then theactivated end of the chain would be removed, and synthesis would stop.Īnswer 5.5. Chain synthesis would occur in a 3’ to 5’ direction. All these steps are similar to those inthe tail-growth mechanism at the 3’ end, except that the nonactivated end ofthe incoming nucleotide initiates the reaction with the activated end of thegrowing chain. The b- and g-phosphateswould be liberated as pyrophosphate. The 3’ hydroxylon an incoming nucleotide could react with the a-phosphate of the 5’ nucleotide by a nucleophilic attack. A hypothetical head-growth mechanismfor DNA synthesis would have the 5’ end of the primer at the active site this5’ end would have a triphosphate on the last nucletide added. 5.7 you would not expect to see a slow-sedimenting peakof nascent DNA.Īnswer5.4. Therefore, by the model of semidiscontinuoussynthesis shown in Fig. Because the short Okazaki fragments shouldstill be in duplex with the large parental DNA strands, the duplex would notseparate from the bulk of the DNA. In an neutral sucrose gradient, the two strands ofthe DNA duplex should stay together. In contrast to the replication eyes, the two new strands are notsynthesized simultaneously at the replication fork in D loop replication.Īnswer 5.3. 
Theproduction of LL shows that replication is not random.Īnswer5.2. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
